Within the scope of its activities related to maritime affairs the Joint Research Centre participates in research addressing fish population biology. The analysis and characterisation of fish populations is of great scientific interest and has implications for conservation and fisheries management. REMUS was a collaborative research project with Prof. R. Hanel from IFM-GEOMAR (Leibniz Institute for Marine Science, University of Kiel), aiming at the identification and characterisation of Mullus surmuletus populations in European waters. The results of this research along with documentary material is freely accessible and should provide information and technical support to the scientific community, but also other stakeholders.
Goatfishes of the family Mullidae show a worldwide distribution in tropical to temperate waters and are of considerable economic importance. Especially in the north-eastern Atlantic and the Mediterranean Sea, red mullet (Mullus barbatus) and striped red mullet (Mullus surmuletus) are major targets for small scale and semi-industrial fisheries, leading to heavy exploitation of populations (Renones et al., 1995).
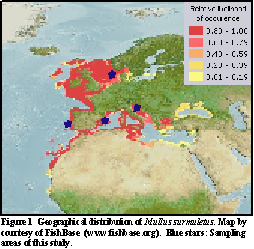
On the European scale Mullus surmuletus shows a wide distribution ranging from the Eastern Atlantic over the North Sea through the English Channel south to Dakar, Senegal and the Canary Islands, including the Mediterranean and the Black Sea (Figure 1).
It is generally considered a warm-temperate Lusitanian species and interestingly a recently recorded increase in abundance observed in commercial catch data of this species in the northern North Sea has been explained by rising water temperatures due to climate change (Cook and Heat, 2005). While striped red mullet has originally been described as a typical sand-dwelling carnivorous species living at depths of less than 100 m (Labropoulou and Eleftheriou, 1997) others authors more differentially point out its competitive adaptation to coarse grain habitats at shallower depths (Lombarte, 2000).
Although there has been considerable research on various aspects of life history traits and habitats of Mullus surmuletus e.g. (Golani, 1994), knowledge on its genetic population structure is rather scarce. Population genetic analyses based on allozyme variation (Mamuris et al., 1999; Mamuris et al., 1998), random amplified polymorphic DNA (Mamuris et al., 1999; Mamuris et al., 1998) and RFLPs of mitochondrial DNA (Apostolidis, 2001) revealed a high degree of polymorphism but low genetic structuring. To further investigate the population of structure of the striped red mullet, specimen of four different sampling areas (Figure 1) have been analysed genetically, by sequencing parts of the mitochondrial genome. Additionally parasite infestation has been examined and morphological data recorded.
Cited Literature
[1] Renones, O., Massuti, E., Morales-Nin, B. (1995) Life history of the red mullet Mullus surmuletus from the bottom-trawl fishery off the Island of Majorca (north-west Mediterranean). Marine Biology, 123 (3), pp. 411-419.
[2] Cook, R.M. and Heath, M.R. (2005) The implications of warming climate for the management of North Sea demersal fisheries. ICES Journal of Marine Science, 62 (7), pp. 1322-1326.
[3] Labropoulou, M. and Eleftheriou, A. (1997) The foraging ecology of two pairs of congeneric demersal fish species: Importance of morphological characteristics in prey selection. Journal of Fish Biology, 50 (2), pp. 324-340.
[4] Lombarte, A., Recasens, L., Gonzalez, M. and De Sola, L.G. (2000) Spatial segregation of two species of Mullidae (Mullus surmuletus and M. barbatus) in relation to habitat. Marine Ecology Progress Series, 206, pp. 239-249.
[5] Golani, D. (1994) Niche separation between colonizing and indigenous goatfish (Mullidae) along the Mediterranean coast of Israel. Journal of Fish Biology, 45 (3), pp. 503-513.
[6] Mamuris, Z., Stamatis, C. and Triantaphyllidis, C. (1999) Intraspecific genetic variation of striped red mullet (Mullus surmuletus L.) in the Mediterranean Sea assessed by allozyme and random amplified polymorphic DNA (RAPD) analysis. Heredity, 83 (1), pp. 30-38.
[7] Mamuris, Z., Apostolidis, A.P., Theodorou, A.J. and Triantaphyllidis, C. (1998) Application of random amplified polymorphic DNA (RAPD) markers to evaluate intraspecific genetic variation in red mullet (Mullus barbatus). Marine Biology, 132 (2), pp. 171-178.
[8] Apostolidis, A.P., Mamuris, Z., Karkavelia, E. and Alifakiotis, T. (2001) Comparison of Greek breeds of horses using RAPD markers. Journal of Animal Breeding and Genetics, 118 (1), pp. 47-56.